Working at home last Thursday, trying to get some quiet time to focus on a review article, I caught the lunchtime news and heard of the untimely death of the actress Wendy Richard. She was best known as Pauline Fowler from long-running BBC soap Eastenders. But those of an—ahem—older generation may remember her more fondly as the ditzy Miss Brahms, one of the bizarre cast of characters from Are you being served?, a 1970s sitcom about the staff of Grace Brothers Department store. Despite the title, service seemed to be the least of the their preoccupations.

Wendy Richards as Miss Brahms
Which is a bit like Open Access sometimes, a topic that crops up frequently, most recently on Caryn Schectman’s blog. It’s supposed to be about serving information more freely to the scientific community (and anyone else who’s interested) but sometimes, although the data are now public (_The truth is out there!_), they’re not always so accessible. I’m not talking about papers—download, print (or perhaps not), read—but all the raw data that are now accumulating at sites around the world. As Heather has mentioned, getting your hands on it is not always that easy.
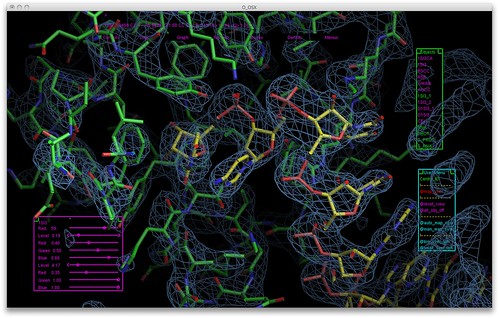
But returning to my desk last Thursday, I found a very nice example of Open Access working well—for me at least. I wanted to check out a detail in a figure of a protein-RNA complex in one of the papers I was reading. Was that hydroxyl group really in that position I asked myself. I wanted to to have a look at the electron density map that the model had been built into for myself.
And I could, using the Electron Density Server (EDS). Let me explain very briefly: when protein crystallographers publish a new structure these days, they are required to deposit not only the coordinates of the macromolecule with the PDB, the Protein Data Bank, so that anyone can have a look at it on their computer*, but they must also send in the structure factors, colloquially known as F’s. The F’s are basically the raw data and allow you to reconstruct the electron density map that was used to guide construction of the atomic model. But only a seasoned crystallographer would know what to do with them; and even then, there is a bit of an activation barrier between grabbing the F’s and running the programs needed to generate the maps.
Thanks to the Electron Density Server, it took me all of 30 seconds to go from “Hmm, I’d like to check that out” to “Oh, I see what they mean.” On the EDS web-page you simply enter the PDB identifier for the structure (taken from the paper) and it immediately serves up a package of files that, once unzipped, lets you fire up the molecular graphics program O. You can then get straight to work: in the O session the structure coordinates and maps are already loaded. The EDS is a fantastic piece of work.
Now, it’s still not for everyone. In part that’s because of the nature of the data, since you need to have some understanding of the crystallographic method to be able to interpret the maps. And O does have a bit of a learning curve (though if you can find a friendly local crystallographer, I reckon they could take you through the basics in 10-20 minutes). So, if you’re a neophyte, you would probably want pretty badly to have a look before you had a go. But for the crystallographic community the EDS is a great resource – it serves up the data with a smile.
Now I’m still not ready to run with the radicals of the Open Science movement who want to lay every single pipette action before the public in real time, but—if a result is published—I do want to be able to see the data. The EDS is, I hope, part of a growing movement that is affecting all fields of science. So I look forward to easier access, in all areas.
*Though it’s also an interesting question of how easy it is for non-structural biologists to grapple with pdb files and molecular graphics programs, even when they don’t care about the electron density. Perhaps a topic for another day.
Gerard J. Kleywegt, Mark R. Harris, Jin yu Zou, Thomas C. Taylor, Anders Wählby, T. Alwyn Jones (2004). The Uppsala Electron-Density Server Acta Crystallographica Section D Biological Crystallography, 60 (12), 2240-2249 DOI: 10.1107/S0907444904013253





Stephen – Once when talking to a group of publishers and librarians I used a similar analogy. I talked about the dissatisfaction of some in the academic community with the way scientific publishing had developed and said they asked “Are we being served?”. And then along comes the OA movement and says “I’m free”.
Har, Har Frank – nice one!
Of course, it’s only free at the point of use. Generating and hosting the content is not gratis. And, as I tried to make out, it has to be served in an easily digestible manner.
The weight of those false eyelashes must have been crippling.
She was young and fit at the time.
Do you know the series? I was surprised to see old re-runs on public TV when I was in Boston in the ’90s – seemed to be very popular with some in the US.
I think I saw a couple of episodes once in my dim and distant youth – I have a vague memory of being at a friend’s house (we were “not allowed” TV) and seeing it in B&W.
I think the data distinction seems a good one – if all the journals asked their authors to submit data in a public dbase upon publication, then this would be a lot easier for everyone to search. Some, but not all, journals already do this. I also think that “negative” results, ie those experiements that are done but are not “used” for anything, could be put somewhere in an online archive… several of which now exist for the purpose.
A complete non-sequitur – my husband yesterday went to Biophysics – he phoned tonight and I overheard my younger daughter saying “have you solved muscle contraction yet?” (I think she always says something like this when he’s away and phones her, to show him she’s interested and has noticed he’s gone – but she’s failed to distinguish between a conference and an experimental trip.)
I watched it myself in the 70s pretty regularly as I recall. Was a bit horrified when I saw i again much later.
On the subject of data deposition, it is complicated by the fact that there are so many different formats. It’s a bit mind-boggling!
Though I would like to see journals getting together to standardise formats for Supplementary Information: I think it should be as a pdf with text and figures integrated in a way that is closer to the format of a published paper than to a submitted manuscript. That’s too much of a challenge for some authors, I guess. But at the very least, figures and legends should be presented on the same page.
Oh, and, at least your daughter is trying to sound interested!
Oh, I watched ‘Are you being served?’ back in the 70s myself – it was a very funny show. I wonder how funny I would find it today…
Do you have to use O? I love O, but the new generation (feh) find it a bit ‘difficult’. They’re brought on Coot, which is nice, but if you’re going to do that you should go the whole hog and have iStructure.
Stephen – I have to say that your segue between Are You Being Served and open-access publishing was shockingly abrupt. I could see the join, mate.
I watched Are You Being Served in the 80s. Were those reruns?
[internets]
No! I remember an episode about a hideaway bed, and Wikipedia says that was aired in 1985 for the first time (would have been in Holland a few years later, with subtitles and stuff). Amazing. My memory is a storage place of insignificant details…
It’s exactly the start of this fragment that I remembered. I was about eight I guess, when I saw it, and did not get any of the innuendo =)
@Richard – the package is specifically set up to allow you to start running O. If you do that in the directory where everything’s been downloaded, then the models and maps are loaded automatically. It’s no surprise that O is the preferred destination since the guys who set up the EDS were largely responsible for O (Alwyn Jones and friends). You could probably load the same files into PyMOL or coot but you’d have to concoct some sort of script that asks for the PDB identifier and then imports the file into that program.
You swine, I acutally looked for iStructure! If only we could get Jobs on the job though, eh?
@Henry – your eyesight is truly exceptional. But I’m afraid that’s the disjointed way my mind works…!
@Eva – thanks for the clip. 1985 was the last year of original broadcasts – the show started in 1972 (according to my copy of Wikipedia at any rate). I remember seeing it at home n the 70s when I was too young to get any of the innuendo (apart perhaps from Mr Humphrys…) – never really understood why Mrs Slocombe kept mentioning her cat!
Wow, I hadn’t even heard of the EDS. I am going to have to play around loading the structures into coot. The EDS set up for O is another classic example of file incompatibility that plagues macro crystallography.
Yes the lack of integration can be a pain. Would certainly be interested to know how you get on (even though I’m a bit old school and haven’t yet embraced coot).
Oh, and welcome to NN Ted – I gather you’ve only recently joined us. Nice to have another crystallographer in town! I saw your very interesting-looking website and will be looking forward to future postings.
Thanks for the welcome. I just posted a piece on how to use the EDS (thanks to you) and it works like a charm with Coot.
Cheers Ted- that was a nice post. As a gesture of thanks I’ll fix the link to your website that I somehow screwed up in my comment above.
By the way, I tried to leave a comment on your site but it decided somehow that I was a ‘spambot’ – not sure how it managed to figure that one out. The people round here still haven’t noticed…
Thanks for heads up on the spambot notification – I just tried to fix it. I am a blogging newbie of 2 weeks so am still learning the ropes.
That seemed to work Ted – I just left a comment and it seemed to be accepted (just waiting for ‘moderation’ – you are playing it safe!).
Hi,
I saw many of you mention AYBS here, so I thought this might interest you.
I run an official “Are You Being Served?” site at:
http://www.aybscentral.com
There are a lot of images and info pertaining to all the cast members.
Enjoy!
Elina